Landscape of circulating diagnostic biomarkers in pancreatic malignancies
Introduction
Pancreatic cancer, the bulk of which is pancreatic ductal adenocarcinoma (PDAC), has very poor prognosis, as more than 50% of cases are diagnosed at an advanced stage (1). Pancreatic cancer has several known risk factors, including smoking, long-standing diabetes, chronic pancreatitis, and obesity (2). Even among patients with known genetic predisposition for PDAC (i.e., mutations in BRCA1, BRCA2, p53, LKB1-STK11, MSH genes), there are no robust biomarkers to aid in screening for the development of pancreatic malignancy.
Retrospective studies have shown that the most common presenting symptoms in PDAC include fatigue and weakness, weight loss, abdominal pain, or the development of jaundice particularly in patients whose tumor is localized to the pancreatic head (3). In the minority (15–20%) of patients with disease localized to the pancreas without significant vascular involvement, surgical resection can prove curative. Many patients also undergo adjuvant treatment with chemotherapy (usually a FOLFOX or gemcitabine based regimen) with or without radiation therapy. Despite available therapies, the 5-year survival for these patients stands at less than 10% (4).
Given the significant morbidity and mortality associated with PDAC, there is a need for the development of robust biomarkers to use as screening tools in detecting asymptomatic premalignant or localized tumors amenable to early intervention. Despite this mortality, PDAC only represents 2.5% of all cancers, necessitating the need for a highly specific and sensitive test to avoid false positives that can prove economically and emotionally burdensome given follow-up diagnostic imaging for individuals and healthcare systems alike (5). Sequencing analyses to evaluate the clonal relationships between primary and metastatic PDAC have shown that nearly five years elapse between the first non-metastatic malignant cell and the emergence of distant disease, suggesting a window for possible clinical screening and intervention in selected patients that may improve outcomes (6).
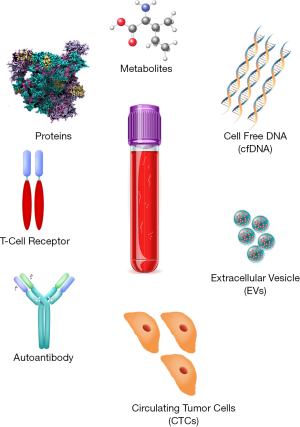
In this review, we explore the landscape of available and emerging blood based biomarkers in the diagnosis of PDAC (Figure 1).
Protein biomarkers, auto-antibodies, and T-cell receptors
Carbohydrate antigen 19-9 (CA 19-9) is among the most widely used biomarkers to aid in the diagnosis of PDAC. A recent meta-analysis of about 2,000 patients to evaluate the utility of CA 19-9 in pancreatic cancer indicated a pooled sensitivity and specificity of 0.8, and an AUC of 0.87 (7). However, it is key to note that these patients were predominantly symptomatic, and CA 19-9 is not a particularly good screening tool in asymptomatic patients (positive predictive value 0.5–0.9% (8).
Moreover, since CA 19-9 is an epitope of sialylated Lewis group antigen, about 5–10% of Caucasian patients lack this genotype and could have a false-negative test (9). CA 19-9 can also be elevated in many benign gastrointestinal conditions as well as other malignancies, including pancreatitis, cirrhosis, cholangitis, and colorectal cancers, which can further complicate accurate diagnosis of pancreatic malignancy in the undifferentiated patient.
Carcinoembryonic antigen (CEA) is a glycoprotein that is also known to be elevated across several malignancies, including in about 30–60% of pancreatic cancer patients. Meta-analyses examining the value of CEA (~3,600 patients across 19 studies) indicated a pooled sensitivity of 0.43 and a specificity of 0.82, which is slightly worse than those of CA 19-9 (10).
Given these analyses, there has also been interest in using combinations of biomarkers isolated from serum, with the hope that panel-based approaches might increase the positive predictive value in early identification of pancreatic cancer. For instance, one study examined over 80 different serum proteins across pancreatic cancer patients, and demonstrated that a panel of CA19-9, ICAM-1, and osteoprotegerin (OPG) enhanced the sensitivity and specificity of discerning malignancy from healthy controls in a validation cohort compared to CA19-9 alone (11). Still another study examined combining CA19-9 with cathepsin D and MMP-7 similarly improved the sensitivity compared to CA19-9 alone (88% vs. 74%) (12).
In recent years, the interplay between evasion of immune destruction and the onset of malignancy has been increasingly appreciated, and it is conceivable that markers of immune modulation may be detected prior to onset of clinical disease (13). Aberrant tumor-specific mutations are now known to result in neoantigens that could be immunogenic and detected by the immune system, and can also result in auto-antibodies that can be detected in sera (14). For instance, MUC1 is known to be overexpressed in PDAC, and an anti-MUC1 antibody assays showed a sensitivity and specificity of 77% and 95%, respectively, in discriminating pancreatic cancer from pancreatitis (15,16). Along this vein, identification of auto-antibodies to overexpressed or mutated proteins in PDAC can aid in early detection. Indeed, studies have also identified T-cell reactivity with specific MUC16 neoantigens in PDAC to correlate with long-term survival in pancreatic cancer (>5 years), suggesting that this approach may be both diagnostic and prognostic in PDAC (17).
The immune microenvironment surrounding the tumor is also often rich with inflammatory cells (i.e., macrophages, regulatory T-cells, myeloid-derived suppressor cells) that secrete various cytokines that can be detected in sera. A prior systematic review compiling studies across 41 cytokines indicated that individual cytokines often exhibit poor diagnostic performance (sensitivity and specificity ~ 50–90%), but showed that six cytokines (IL-1β; IL-6, IL-8, VEGF, TGF, IL-10) were consistently correlated with PDAC (18). The levels of these cytokines in sera have been correlated with PDAC stage (higher levels implying poor patient prognosis), but the principal limitation is that these cytokines are commonly elevated across many malignancies and many other benign conditions where systemic inflammation is present.
These combination approaches do represent a promising strategy in enhancing the AUC and sensitivity/specificity for PDAC, and are actively being explored. However, the limited specificity of these proteins to cancer, the lack of consistent protein biomarker utility in early cancer detection, and the variability of multi-parameter protein assays dependent on antibody performance remain key barriers in the success of purely protein-based assays. In addition, many of these studies have used healthy controls that may not be age- or comorbidity-matched, and can prove challenging to apply in the clinical setting without larger scale validation (19). However, some of these biomarkers are being developed for molecular imaging of PanIN lesions using sophisticated liposomal and radiolabeled targets. The ability to localize protein biomarkers with imaging is necessary to advance early detection strategies to the point of intervention.
Metabolites
It is well established that pancreatic cancer cells undergo significant metabolic pathway modification to enhance survival in hypoxic and nutrient poor environments (20). The substrates, intermediates, and by-products of these aberrant metabolic pathways can be quantified, and may be useful in developing metabolomics signatures of early stage pancreatic cancer (21). Specifically, in PDAC, the near-universal mutations in KRAS lead to activation of the MEK/ERK/PI3K/AKT pathways, and cause cells to engage in autophagy and micropinocytosis to provide substrates for accelerated metabolic demand (22,23).
Preclinical models examining the metabolic landscape of pancreatic intraepithelial neoplasia (PanIN), an early stage of the disease, to PDAC using NMR spectroscopy revealed significant differences in kynurenate and methionine levels (24). These levels were elevated in PanIN but decreased in PDAC, likely because of alterations in methionine and tryptophan metabolism from upregulation of specific amino-acid transporters that allows for disease progression and metastasis. PanINs are too small to discern from non-tumor regions on conventional imaging, but these metabolomics approaches may aid in detection of early-stage disease if similar approaches are validated in human trials. A separate large (914 subject) cohort showed that blood derived metabolite biomarkers can successfully distinguish between chronic pancreatitis and pancreatic cancer (25). Using GC-MS and LC-MS metabolomics techniques, these authors identified a signature of 9 metabolites (i.e., sphingolipids, ceramides) and CA19-9 that showed an NPV of >99% in the training set and >99% in the validation set.
Another approach profiling metabolites in plasma from pancreatic cancer patients and matched controls indicated that patients had elevated branched-chain amino acids (Ile, Leu, Val) up to 5 years prior to PDAC diagnosis, corresponding to the 2× increased risk (26). These results suggest that protein breakdown occurs much earlier than clinical overt evidence of cachexia that PDAC patients often present with. Follow-up murine studies showed that these findings are driven by Kras mutations, validating the notion that alterations in metabolic pathways driven by genetic predispositions can be quantified and prove diagnostic in early detection of PDAC.
Cell-free DNA (cfDNA) and RNA
cfDNA is non-encapsulated DNA found in the bloodstream, released by apoptosis and cell necrosis. Cell-free DNA released from tumor cells is referred to as circulating-tumor DNA (ctDNA) or tumor-derived cfDNA. Advances in next-generation sequencing now allow for characterization of ctDNA to identify mutations and serve as a ‘liquid biopsy’ for early detection of pancreatic malignancies. Indeed, a study examining a cohort of 640 patients with various malignancies found ctDNA to be detected in sera in >75% of cases. More specifically, its sensitivity and specificity in identification of clinically relevant KRAS mutations was 87% and 99% respectively (27). In addition, there was a greater than 90% concordance between the mutations identified in cfDNA and those identified from the primary tumor biopsy.
A separate study examining 26 patients with pancreas and biliary malignancies showed similar results, where 90% of the mutations noted in tumor biopsies were also seen in cfDNA (28). A more recent approach termed CancerSEEK combining mutations in cfDNA with protein tumor biomarkers examined 1,005 patients with localized cancers of the lungs, gastrointestinal, or genitourinary tract (29). Their study showed sensitivities between 69–98%, and sensitivities >99% for detection of cancers for which there are no screening tests (i.e., ovarian, pancreas).
Several additional genetic polymorphisms and mutations have also been implicated in the development of PDAC, and could help facilitating screening or be corroborative in diagnosis (30). For example, a recent study examining about 2,900 PDAC cases alongside over 5000 controls demonstrated that BRCA2 mutation K3326X and CHEK2 mutation I157T were associated with an increased risk (~1.8×) of developing PDAC (31). Similarly, the telomerase (TERT) gene locus, as well as caspase-9 and survivin gene polymorphisms have been implicated in pancreatic cancer development (32,33). Although several of these pathways are known to be involved in oncogenesis, the exact mechanism remains to be elucidated.
Long non-coding RNA (lncRNA) and micro RNAs have also been implicated in the pathogenesis of pancreatic cancer. LncRNAs in particular demonstrates high molecular stability, and has been shown to be detectable in plasma (34). For example, higher expression of lncRNA MALAT1 has been shown to correlate with poorer PDAC survival, likely though MALAT1 mediated upregulation of genes involved in EMT (35). Several microRNAs have also been associated with PDAC (i.e., miR-21 and miR-155), and correlate with tumor stage or prognosis (36). Additional studies are required to tease out the exact post-transcriptional regulatory mechanisms involved with different miRNAs to determine why they correlate with disease progression.
Beyond cfDNA mutation analysis, studies have also examined using cfDNA epigenetic markers to further refine diagnostic sensitivity and specificity. One study examining a cohort of patients with chronic pancreatitis versus pancreatic cancer studied the methylation status across a set of gene promoters from cfDNA (37). Their results showed a 91% sensitivity and 90% specificity in distinguishing malignancy from more benign inflammatory disorders. A more recent study demonstrated the ability of methylation profiles of cfDNA to discriminate between patients with different malignancies (38). In this work, 24 early stage PDAC and 24 healthy control methylation cfDNA profiles were used to demonstrate the ability to use differentially methylated regions (DMRs) to detect early cancer. They then developed DMR in a discovery cohort of 189 plasma samples from a broad range of cancers including PDAC, colorectal cancer, breast, cancer, lung cancer, renal cancer, bladder cancer and acute myeloid leukemia. Testing in a validation cohort of 199 samples of PDAC, lung cancer, leukemia, and healthy controls yielded an area under the receiver operator characteristic curve of 0.918 for PDAC detection. Altogether, cfDNA methylation profiles provide additional information to mutational analysis.
Taken together, these studies demonstrate that detection of malignancies is possible with high sensitivity and specificity without a priori knowledge of the tumor genotype. However, cfDNA KRAS mutation detection alone is limited in the ability to determine the tumor site of origin and there are a minority of PDAC cases that lack classic KRAS mutations leading to possible false-negatives. The combination of cfDNA genotyping with methylation profiling and protein based biomarkers appears to be a robust strategy to boost diagnostic accuracy.
Extracellular vesicles (EVs)
EVs are lipid-bilayer enclosed vesicles that are released from the surface of plasma membranes and are thought to facilitate cell-cell communication. Exosomes are a population of EVs that are 40–120 nm in size that are the most common EVs used in studies. They have the potential to transmit cargo, including DNA, mRNA, miRNA, and proteins between cells and have been implicated in tumor progression, invasion, immune tolerance, and drug-resistance (39).
In the context of PDAC, studies have demonstrated that tumor-derived EVs are taken up by cells in the liver, thereby causing upregulation of TGF-beta and uptake of macrophage migration inhibitory factor (40). This enables the creation of a fibrotic microenvironment that facilitates the metastasis of pancreatic cancer to the liver. Indeed, blockade of MIF transmission prevented liver metastasis, and MIF was elevated in PDAC patients who progressed to liver metastasis compared to those who did not progress. Another study indicated that pancreatic cancer derived EVs carrying a specific miRNA (miR-203) downregulated innate immune responses and cytokines (TNFa, IL-12) involved in dendritic cell activation, likely helping the tumor evade immune detection (41).
Therefore, understanding the composition and function of PDAC-derived EVs can provide clues in the events leading to PDAC progression and can aid in early diagnosis. A key challenge in using exosomes is in identifying cancer-specific exosomes from those made by healthy cells, and specific markers are required to eliminate this confounding effect. As proof-of-concept, one study showed that PDAC-derived exosomes were enriched in glypican-1 (GPC1, a cell surface proteoglycan), and also showed that these EVs carried mutant KRAS DNA (42). Their analysis indicated that GPC1 EVs could be detected in both precursor lesions like PanIN and pancreatic cancer, and could distinguish between any evidence of malignancy and healthy patients with an AUC of 1 (100% sensitivity, 100% specificity). Although subsequent studies have not been able to replicate the performance of GPC1+ EVs, these studies have demonstrated the utility of EV content as a strategy to enhance sensitivity and specificity for blood based biomarkers (43,44).
Two additional studies have also attempted to use EV DNA to identify KRAS mutations known to drive PDAC. KRAS G12D mutations were identified in 7.4% of control patients, 67% of localized PDAC, 80% of locally advanced PDAC, and 85% of metastatic PDAC patients (45). A separate study showed that KRAS G12D was seen in EV DNA in nearly 40% of patients with PDAC, 29% of patients with IPMN, and only 3% in controls (46).
A study with miRNA isolated from EVs revealed a cocktail of miRNAs (miR-1246,4644,3976,4306) upregulated in 83% of pancreatic cancer derived EVs, but not in control groups (47). Measurement of these specific miRNAs in combination with a protein panel (CD44v6, Tspan8, EpCAM, MET, and CD104) allowed for distinction between PDAC and non-PDAC controls with sensitivity 100% and specificity 80%. The control group in this study had also included cases of chronic pancreatitis and benign pancreatic lesions.
There are several advantages in using exosome-based strategies for early PDAC detection, namely that exosomes are stable even in frozen plasma or serum derived from patients, exhibit high sensitivity as shown above, and contain heterogeneous cargo (DNA, RNA, proteins) providing a multi-prong approach towards distinguishing PDAC from healthy patients.
Circulating tumor cells (CTCs)
CTCs are cells derived from the primary that have entered into the bloodstream and may seed a distant metastatic site (48). CTCs are low-frequency, occurring in 1 per billion cells in patients with malignancies, and therefore present challenges in isolation for characterization and further study (49).
In PDAC, several studies have demonstrated isolation of CTCs regardless of stage among localized, locally advanced, or metastatic patients (50-52). For instance, a prospective study of 100 PDAC patients prior to treatment indicated a sensitivity of 75%, specificity of 96%, and 100%, AUC of 0.867 concordance of Kras mutation subtype between the primary tumor and CTCs (53). Interestingly, the presence of >3 CTCs per 4mL of blood was able to distinguish between locoregional and metastatic disease.
These data suggest that early stage PDAC patients would show even rarer CTCs in circulating sera compared to those with advanced disease, and as such, methods development to better isolate and characterize CTCs has been of keen interest. Microfluidic devices chemically functionalized with anti-EPCAM (epithelial-cell-adhesion-molecule) provide specificity for CTC capture as EPCAM is overexpressed in CTCs but absent in hematologic cells (54). Given this methodology, both flow velocity and shear force are key parameters in ensuring that CTCs can be successfully captured for study. Molecular analysis of CTCs has also yielded insights into PDAC biology, and one study identified Wnt2 to be enriched in CTC populations, suggesting its role in promoting anchorage-independence and metastasis (55). However, one key limitation of EPCAM-based enrichment is that it may miss malignant cells that have undergone epithelial to mesenchymal transition.
More recent studies utilizing a negative depletion microfluidic platform (to reduce WBC contaminants) demonstrated that it was possible to collect CTCs from precursor lesions like IPMN. This platform called the CTC-iChip avoids the previous issues of EPCAM enrichment bias from previous CTC technologies. In this study, RNA-sequencing analysis of MUC genes confirmed that the origin of CTCs was from the pancreas, thereby increasing the window of sensitivity in diagnosing earlier stage disease (56). Given these promising results, there is a need for large-scale validation of these results in human trials prior to clinical adoption.
Conclusions
The majority of patients diagnosed with pancreatic cancer present with advanced disease, and succumb to their disease within a few years of diagnosis. Screening methodologies that enable earlier detection of PDAC hold significant promise in directing patients towards curative surgical therapies. Among circulating biomarkers, many options exist, including development of proteomic-panels, discerning immunologic and cytokine signatures of PDAC, identifying mutations or methylation changes in cfDNA, and isolating EVs or CTCs in moving towards early detection. Development of a test with optimal sensitivity and specificity will likely involve a combination of these approaches, and active collaboration will be required to coordinate future prospective trials in high-risk population to validate assays prior to widespread adoption.
Acknowledgments
Funding: None.
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/apc.2020.03.04). DTT has received consulting fees from Merrimack Pharmaceuticals, Ventana Roche, Foundation Medicine, Inc., and EMD Millipore Sigma, which are not related to this work. DTT receives research support from ACD-Biotechne, which was not used in this work. DTT is a founder and has equity in ROME therapeutics, PanTher Therapeutics and TellBio, Inc. TellBio, Inc. is a company developing circulating tumor cell diagnostics and therapeutics that is relevant to this article, but no funding was provided by the company for this work. DTT has intellectual property that has pending licensing to TellBio, Inc. YGHB reports personal fees from Public Analysis Inc (for Takeda). DTT’s interests were reviewed and are managed by Massachusetts General Hospital and Partners Health Care in accordance with their conflict of interest policies. VS has no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin 2019;69:7-34. [Crossref] [PubMed]
- Ryan DP, Hong TS, Bardeesy N. Pancreatic adenocarcinoma. N Engl J Med 2014;371:1039-49. [Crossref] [PubMed]
- Porta M, Fabregat X, Malats N, et al. Exocrine pancreatic cancer: symptoms at presentation and their relation to tumour site and stage. Clin Transl Oncol 2005;7:189-97. [Crossref] [PubMed]
- Moutinho-Ribeiro P, Macedo G, Melo SA. Pancreatic Cancer Diagnosis and Management: Has the Time Come to Prick the Bubble? Front Endocrinol (Lausanne) 2019;9:779. [Crossref] [PubMed]
- Rawla P, Sunkara T, Gaduputi V. Epidemiology of Pancreatic Cancer: Global Trends, Etiology and Risk Factors. World J Oncol 2019;10:10-27. [Crossref] [PubMed]
- Yachida S, Jones S, Bozic I, et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 2010;467:1114-7. [Crossref] [PubMed]
- Huang Z, Liu F. Diagnostic value of serum carbohydrate antigen 19-9 in pancreatic cancer: a meta-analysis. Tumour Biol 2014;35:7459-65. [Crossref] [PubMed]
- Ballehaninna UK, Chamberlain RS. The clinical utility of serum CA 19-9 in the diagnosis, prognosis and management of pancreatic adenocarcinoma: An evidence based appraisal. J Gastrointest Oncol 2012;3:105-19. [PubMed]
- Goggins M. Molecular markers of early pancreatic cancer. J Clin Oncol 2005;23:4524-31. [Crossref] [PubMed]
- Meng Q, Shi S, Liang C, et al. Diagnostic and prognostic value of carcinoembryonic antigen in pancreatic cancer: a systematic review and meta-analysis. Onco Targets Ther 2017;10:4591-8. [Crossref] [PubMed]
- Brand RE, Nolen BM, Zeh HJ, et al. Serum biomarker panels for the detection of pancreatic cancer. Clin Cancer Res 2011;17:805-16. [Crossref] [PubMed]
- Park HD, Kang ES, Kim JW, et al. Serum CA19-9, cathepsin D, and matrix metalloproteinase-7 as a diagnostic panel for pancreatic ductal adenocarcinoma. Proteomics 2012;12:3590-7. [Crossref] [PubMed]
- Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell 2011;144:646-74. [Crossref] [PubMed]
- Schumacher TN, Schreiber RD. Neoantigens in cancer immunotherapy. Science 2015;348:69-74. [Crossref] [PubMed]
- Hinoda Y, Ikematsu Y, Horinochi M, et al. Increased expression of MUC1 in advanced pancreatic cancer. J Gastroenterol 2003;38:1162-6. [Crossref] [PubMed]
- Gold DV, Modrak DE, Ying Z, et al. New MUC1 serum immunoassay differentiates pancreatic cancer from pancreatitis. J Clin Oncol 2006;24:252-8. [Crossref] [PubMed]
- Balachandran VP, Luksza M, Zhao JN, et al. Identification of unique neoantigen qualities in long-term survivors of pancreatic cancer. Nature 2017;551:512-6. [Crossref] [PubMed]
- Yako YY, Kruger D, Smith M, et al. Cytokines as Biomarkers of Pancreatic Ductal Adenocarcinoma: A Systematic Review. PLoS One 2016;11:e0154016 [Crossref] [PubMed]
- England CG, Hernandez R, Eddine SB, et al. Molecular Imaging of Pancreatic Cancer with Antibodies. Mol Pharm 2016;13:8-24. [Crossref] [PubMed]
- Sousa CM, Kimmelman AC. The complex landscape of pancreatic cancer metabolism. Carcinogenesis 2014;35:1441-50. [Crossref] [PubMed]
- Zhou W, Capello M, Fredolini C, et al. Proteomic analysis reveals Warburg effect and anomalous metabolism of glutamine in pancreatic cancer cells. J Proteome Res 2012;11:554-63. [Crossref] [PubMed]
- Almoguera C, Shibata D, Forrester K, et al. Most human carcinomas of the exocrine pancreas contain mutant c-K-ras genes. Cell 1988;53:549-54. [Crossref] [PubMed]
- Pylayeva-Gupta Y, Grabocka E, Bar-Sagi D. RAS oncogenes: weaving a tumorigenic web. Nat Rev Cancer 2011;11:761-74. [Crossref] [PubMed]
- Wen S, Li Z, Feng J, et al. Metabonomic changes from pancreatic intraepithelial neoplasia to pancreatic ductal adenocarcinoma in tissues from rats. Cancer Sci 2016;107:836-45. [Crossref] [PubMed]
- Mayerle J, Kalthoff H, Reszka R, et al. Metabolic biomarker signature to differentiate pancreatic ductal adenocarcinoma from chronic pancreatitis. Gut 2018;67:128-37. [Crossref] [PubMed]
- Mayers JR, Wu C, Clish CB, et al. Elevation of circulating branched-chain amino acids is an early event in human pancreatic adenocarcinoma development. Nat Med 2014;20:1193-8. [Crossref] [PubMed]
- Bettegowda C, Sausen M, Leary RJ, et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med 2014;6:224ra24 [Crossref] [PubMed]
- Zill OA, Greene C, Sebisanovic D, et al. Cell-Free DNA Next-Generation Sequencing in Pancreatobiliary Carcinomas. Cancer Discov 2015;5:1040-8. [Crossref] [PubMed]
- Cohen JD, Li L, Wang Y, et al. Detection and localization of surgically resectable cancers with a multi-analyte blood test. Science 2018;359:926-30. [Crossref] [PubMed]
- Klein AP, Wolpin BM, Risch HA, et al. Genome-wide meta-analysis identifies five new susceptibility loci for pancreatic cancer. Nat Commun 2018;9:556. [Crossref] [PubMed]
- Obazee O, Archibugi L, Andriulli A, et al. Germline BRCA2 K3326X and CHEK2 I157T mutations increase risk for sporadic pancreatic ductal adenocarcinoma. Int J Cancer 2019;145:686-93. [Crossref] [PubMed]
- Campa D, Rizzato C, Stolzenberg-Solomon R, et al. TERT gene harbors multiple variants associated with pancreatic cancer susceptibility. Int J Cancer 2015;137:2175-83. [Crossref] [PubMed]
- Theodoropoulos GE, Michalopoulos NV, Panoussopoulos SG, et al. Effects of caspase-9 and survivin gene polymorphisms in pancreatic cancer risk and tumor characteristics. Pancreas 2010;39:976-80. [Crossref] [PubMed]
- Moschovis D, Gazouli M, Tzouvala M, et al. Long non-coding RNA in pancreatic adenocarcinoma and pancreatic neuroendocrine tumors. Ann Gastroenterol 2017;30:622-8. [PubMed]
- Liu JH, Chen G, Dang YW, et al. Expression and prognostic significance of lncRNA MALAT1 in pancreatic cancer tissues. Asian Pac J Cancer Prev 2014;15:2971-7. [Crossref] [PubMed]
- Papaconstantinou IG, Manta A, Gazouli M, et al. Expression of microRNAs in patients with pancreatic cancer and its prognostic significance. Pancreas 2013;42:67-71. [Crossref] [PubMed]
- Liggett T, Melnikov A, Yi QL, et al. Differential methylation of cell-free circulating DNA among patients with pancreatic cancer versus chronic pancreatitis. Cancer 2010;116:1674-80. [Crossref] [PubMed]
- Shen SY, Singhania R, Fehringer G, et al. Sensitive tumour detection and classification using plasma cell-free DNA methylomes. Nature 2018;563:579-83. [Crossref] [PubMed]
- Mashouri L, Yousefi H, Aref AR, et al. Exosomes: composition, biogenesis, and mechanisms in cancer metastasis and drug resistance. Mol Cancer 2019;18:75. [Crossref] [PubMed]
- Costa-Silva B, Aiello NM, Ocean AJ, et al. Pancreatic cancer exosomes initiate pre-metastatic niche formation in the liver. Nat Cell Biol 2015;17:816-26. [Crossref] [PubMed]
- Zhou M, Chen J, Zhou L, et al. Pancreatic cancer derived exosomes regulate the expression of TLR4 in dendritic cells via miR-203. Cell Immunol 2014;292:65-9. [Crossref] [PubMed]
- Melo SA, Luecke LB, Kahlert C, et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 2015;523:177-82. [Crossref] [PubMed]
- Lai X, Wang M, McElyea SD, et al. A microRNA signature in circulating exosomes is superior to exosomal glypican-1 levels for diagnosing pancreatic cancer. Cancer Lett 2017;393:86-93. [Crossref] [PubMed]
- Yang KS, Im H, Hong S, et al. Multiparametric plasma EV profiling facilitates diagnosis of pancreatic malignancy. Sci Transl Med 2017; [Crossref] [PubMed]
- Allenson K, Castillo J, San Lucas FA, et al. High prevalence of mutant KRAS in circulating exosome-derived DNA from early-stage pancreatic cancer patients. Ann Oncol 2017;28:741-7. [Crossref] [PubMed]
- Yang S, Che SP, Kurywchak P, et al. Detection of mutant KRAS and TP53 DNA in circulating exosomes from healthy individuals and patients with pancreatic cancer. Cancer Biol Ther 2017;18:158-65. [Crossref] [PubMed]
- Madhavan B, Yue S, Galli U, et al. Combined evaluation of a panel of protein and miRNA serum-exosome biomarkers for pancreatic cancer diagnosis increases sensitivity and specificity. Int J Cancer 2015;136:2616-27. [Crossref] [PubMed]
- Gupta GP, Massague J. Cancer metastasis: building a framework. Cell 2006;127:679-95. [Crossref] [PubMed]
- Woo D, Yu M. Circulating tumor cells as "liquid biopsies" to understand cancer metastasis. Transl Res 2018;201:128-35. [Crossref] [PubMed]
- Iwanicki-Caron I, Basile P, Toure E, et al. Usefulness of circulating tumor cell detection in pancreatic adenocarcinoma diagnosis. Am J Gastroenterol 2013;108:152-5. [Crossref] [PubMed]
- Kulemann B, Pitman MB, Liss AS, et al. Circulating tumor cells found in patients with localized and advanced pancreatic cancer. Pancreas 2015;44:547-50. [Crossref] [PubMed]
- Gao Y, Zhu Y, Zhang Z, et al. Clinical significance of pancreatic circulating tumor cells using combined negative enrichment and immunostaining-fluorescence in situ hybridization. J Exp Clin Cancer Res 2016;35:66. [Crossref] [PubMed]
- Ankeny JS, Court CM, Hou S, et al. Circulating tumour cells as a biomarker for diagnosis and staging in pancreatic cancer. Br J Cancer 2016;114:1367-75. [Crossref] [PubMed]
- Nagrath S, Sequist LV, Maheswaran S, et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 2007;450:1235-9. [Crossref] [PubMed]
- Yu M, Ting DT, Stott SL, et al. RNA sequencing of pancreatic circulating tumour cells implicates WNT signalling in metastasis. Nature 2012;487:510-3. [Crossref] [PubMed]
- Franses JW, Basar O, Kadayifci A, et al. Improved Detection of Circulating Epithelial Cells in Patients with Intraductal Papillary Mucinous Neoplasms. Oncologist 2018;23:121-7. [Crossref] [PubMed]
Cite this article as: Sridharan V, Hernandez-Barco YG, Ting DT. Landscape of circulating diagnostic biomarkers in pancreatic malignancies. Ann Pancreat Cancer 2020;3:5.